The Grover Lab
Research
Small RNA Motifs
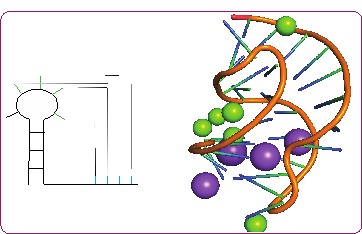
RNA structures are modular and dynamic. Our laboratory studies small structural units that are likely to form in many RNA structures. These motifs are likely to have certain sequence requirements, and associated stability, that are relevant to their function. We are currently investigating the sequence and stability of small RNA in various small loops regions of RNA. We are particularly interested in RNA structures in which metal ions are implicated in structure formation and/or in function.
The RNA we study are often derived from functional regions of large RNA. We often choose RNA that are well characterized. The projects start by designing small RNA and various control RNA and DNA. This requires learning about RNA in general and reading literature on a given RNA. Then we measure the energy it requires to form the structures using methods such as thermal denaturation and isothermal calorimetry. We also use techniques such as fluorescence spectroscopy and native PAGE experiments to understand RNA structures. We are planning new explorations of RNA structures using circular dichroism and Raman spectroscopy.
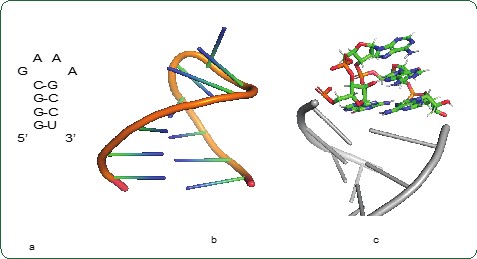
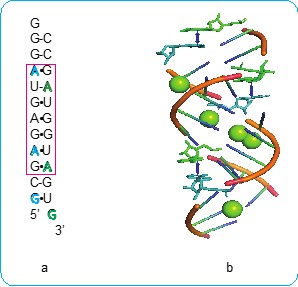
Students who enjoy explorations and are self-motivated to do research will enjoy developing experiments that elucidate the rules of RNA structures. Once the rules of RNA structures are clear, scientists could potentially predict the structure and function of all RNA, including coronavirus. Thus, it is theoretically possible to stop a virus using our knowledge of RNA structures and functions. In the past year, several students have started working on projects related to Coronavirus, some are studying the structures of RNA and others are examining ways to inhibit the coronavirus by designing small RNA-based inhibitors. Along with learning about RNA, students also participate in outreach efforts to share their knowledge about RNA and science with the broader community.
Report an issue -
Last updated: 10/29/2021